data.table join(merge)
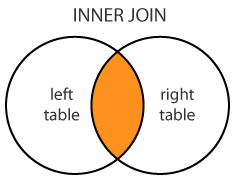
Inner join: merge(x = df1, y = df2, by = "CustomerId")
merge(x = df1, y = df2, by.x="keyx", by.y="keyy")
Cross join: merge(x = df1, y = df2, by = NULL)
Outer join: merge(x = df1, y = df2, by = "CustomerId", all = TRUE)
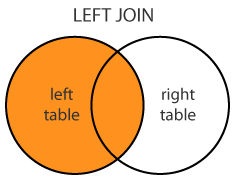
Left outer: merge(x = df1, y = df2, by = "CustomerId", all.x = TRUE)
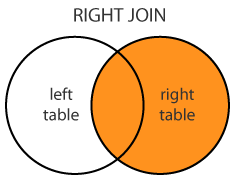
Right outer: merge(x = df1, y = df2, by = "CustomerId", all.y = TRUE)
suffixes=c("x",".y")
data.table의 merge()는 기본모듈인 merge() 과 유사하지만,
data.frame에 기반한 basic merge()는 data.frame내에 변수를 pk로 사용하여 merge하는데 비해
data.table의 Merging은 common key variable 을 pk로 사용하여 merge한다는 점이 다르다.
Key확인 – tables()
setkey는 data.table의 Key로써 A, B를 지정한다. (이 id 변수로 그룹화의 기준으로 생각하면 된다.)
all set* functions change their input by reference
Fast add/ remove/ update subsets of columns, by reference. :=
> key(dt1)
> key(dt2)
[1] "A"
> tables() # KEY column reports the key'd columns
NAME NROW NCOL MB COLS KEY
[1,] dt1 6 2 1 A,X A
[2,] dt2 6 2 1 A,Y A
Total: 2MB
update using Two tables
dt1 <- data.table( A=letters[rep(1:3,2)], X=1:6, key="A") dt2 <- data.table( A=letters[rep(2:4,2)], Y=6:1, key="A") dt1 <- data.table( A=letters[rep(1:3,2)], X=1:6) ; setKey(dt1, "A") # same as above dt2 <- data.table( A=letters[rep(2:4,2)], Y=6:1) ; setKey(dt2, "A")
Inner Join
merge(dt1, dt2, by="A") merge(dt1, dt2) # setKey된 A로 merge dt1[dt2, on=.(A), nomatch=0] # no match row is not returned
A X Y
1: b 2 6
2: b 2 3
3: b 5 6
4: b 5 3
5: c 3 5
6: c 3 2
7: c 6 5
8: c 6 2(Left) OUTER JOIN
merge(dt1, dt2, by="A", all.x = T) dt2[dt1, on="A"]
A X Y A Y X
1: a 1 NA 1: a NA 1
2: a 4 NA 2: a NA 4
3: b 2 6 3: b 6 2
4: b 2 3 4: b 3 2
5: b 5 6 5: b 6 5
6: b 5 3 6: b 3 5
7: c 3 5 7: c 5 3
8: c 3 2 8: c 2 3
9: c 6 5 9: c 5 6
10: c 6 2 10: c 2 6Right (OUTER) JOIN
merge(dt1, dt2, by="A", all.y = T) dt1[dt2, on="A"]
A X Y
1: b 2 6
2: b 2 3
3: b 5 6
4: b 5 3
5: c 3 5
6: c 3 2
7: c 6 5
8: c 6 2
9: d NA 4
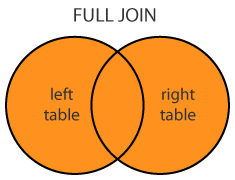
10: d NA 1Full (OUTER) JOIN
merge(dt1, dt2, all=T)
A X Y
1: a 1 NA
2: a 4 NA
3: b 2 6
4: b 2 3
5: b 5 6
6: b 5 3
7: c 3 5
8: c 3 2
9: c 6 5
10: c 6 2
11: d NA 4
12: d NA 1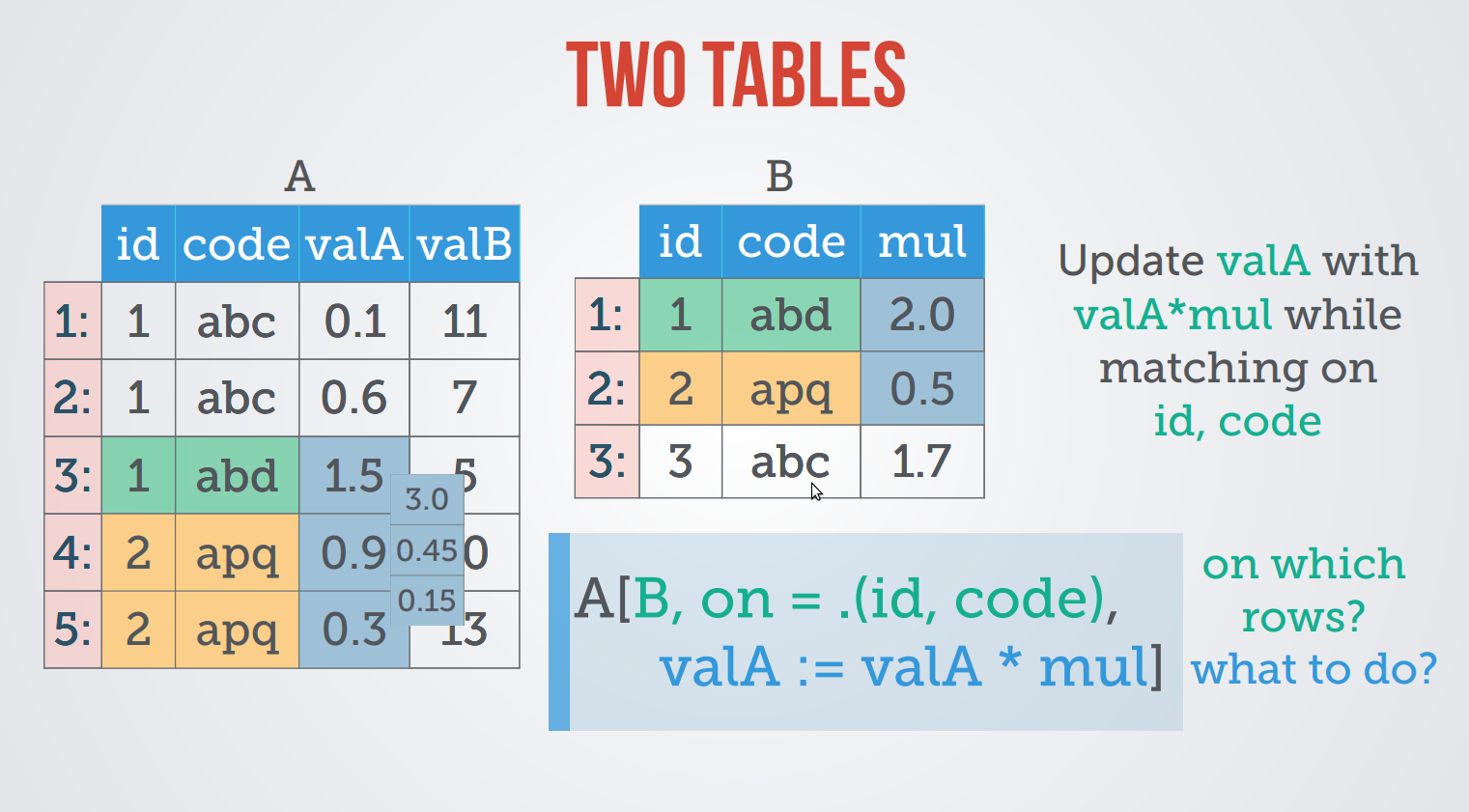

https://github.com/Rdatatable/data.table/wiki/Getting-started
https://stackoverflow.com/questions/1299871/how-to-join-merge-data-frames-inner-outer-left-right
https://stackoverflow.com/questions/8508482/what-does-sd-stand-for-in-data-table-in-r
https://www.analyticsvidhya.com/blog/2016/05/data-table-data-frame-work-large-data-sets/
Use of lapply .SD in data.table R
HTML vignettes: https://github.com/Rdatatable/data.table/wiki/Getting-started
>
>
>
>
>
>
Creates a Join data table
J {data.table}
DT = data.table(x=rep(c("b","a","c"),each=3),
y=c(1,3,6),
v=1:9)
#row selection like join
DT[x=="a"]
DT["a", on="x"]
DT[.("a"), on="x"]
DT[J("a"), on="x"]
setkey(DT, "x")
tables()
DT["a"]
DT[.("a")]
DT[J("a")]
x y v
1: a 1 4
2: a 3 5
3: a 6 6
4: b 1 1
5: b 3 2
6: b 6 3
7: c 1 7
8: c 3 8
9: c 6 9 x y v
1: a 1 4
2: a 3 5
3: a 6 6J : (J)oin.
the same result as calling list. J is a direct alias for list but results in clearer more readable code.
SJ : (S)orted (J)oin.
The same value as J() but additionally setkey() is called on all the columns in the order they were passed in to SJ.
For efficiency, to invoke a binary merge rather than a repeated binary full search for each row of i.
CJ : (C)ross (J)oin.
A data.table is formed from the cross product of the vectors.
For example, 10 ids, and 100 dates, CJ returns a 1000 row table containing all the dates for all the ids.
It gains sorted,
which by default is TRUE for backwards compatibility. FALSE retains input order.
DT = data.table(x=rep(c("b","a","c"),each=3),
y=c(1,3,6),
v=1:9)
#row selection like join
DT[x=="a"]
DT["a", on="x"]
DT[.("a"), on="x"]
DT[J("a"), on="x"]
setkey(DT, "x")
tables()
DT["a"]
DT[.("a")]
DT[J("a")]
Join 시 Match되지 않는 데이터 처리
- 기본적으로 NA로 return된다.
- NA로 처리될 row를 지워버릴수 있다.
- roll을 통해 다른 값을 대체할수 있다.
(LOCF) last observation carried forward => +Inf (or TRUE) 마지막값으로
(NOCB)next observation carried backward => -Inf
rolls the nearest value instead. => "nearest"
DT[.("b", 1:2), on=c("x", "y")] # no match returns NA
x y v
1: b 1 1
2: b 2 NA
DT[.("b", 1:2), on=.(x, y), nomatch=0] # no match row is not returned
x y v
1: b 1 1
DT[.("b", 1:2), on=c("x", "y"), roll=Inf] # locf, nomatch row gets rolled by previous row
x y v
1: b 1 1
2: b 2 1
DT[.("b", 1:2), on=.(x, y), roll=-Inf] # nocb, nomatch row gets rolled by next row
x y v
1: b 1 1
2: b 2 2
Inner join: df1[df2, on=.("CustomerId"), nomatch=0L]
Outer join: merge(x = df1, y = df2, by = "CustomerId", all = TRUE)
Left outer: merge(x = df1, y = df2, by = "CustomerId", all.x = TRUE)
Right outer: dt1[dt2, on=.("CustomerId")]
Cross join: merge(x = df1, y = df2, by = NULL)
dtProd <- data.table(CustomerId = c(1:6), Product= c(rep("iphone",3), rep("gallaxy",3)) )
dtAddr <- data.table(CustomerId = c(3,5,7), Address= c(rep("Seoul",2), rep("Pusan", 1)) )
# full outer join
merge(dtProd, dtAddr, by="CustomerId", all=T)
merge(dtProd, dtAddr, by=c("CustomerId", "Sex"), all=T)
# inner join
merge(dtProd, dtAddr, by="CustomerId")
dtProd[dtAddr, nomatch=0L, on="CustomerId"]
# anti join - use `!` operator
dtProd[!dtAddr, on="CustomerId"]
### outer join
# right outer join (unkeyed)
dtProd %>% setkey(NULL)
dtAddr %>% setkey( NULL)
dtProd[dtAddr, on = "CustomerId"]
# right outer join (keyed data.tables)
dtProd %>% setkey(CustomerId)
dtAddr %>% setkey(CustomerId)
dtProd[dtAddr]
# left outer join - swap dt1 with dt2
dtAddr[dtProd, on = "CustomerId"]
join(merge)
dtProd <- data.table(CustomerId = c(1:6), Product= c(rep("iphone",3), rep("gallaxy",3)) )
dtAddr <- data.table(CustomerId = c(3,5,7), Address= c(rep("Seoul",2), rep("Pusan", 1)) )
# full outer join
merge(dtProd, dtAddr, by="CustomerId", all=T)
merge(dtProd, dtAddr, by=c("CustomerId", "Sex"), all=T)
# inner join
merge(dtProd, dtAddr, by="CustomerId")
dtProd[dtAddr, nomatch=0L, on="CustomerId"]
# anti join - use `!` operator
dtProd[!dtAddr, on="CustomerId"]
### outer join
# right outer join (unkeyed)
dtProd %>% setkey(NULL)
dtAddr %>% setkey( NULL)
dtProd[dtAddr, on = "CustomerId"]
# right outer join (keyed data.tables)
dtProd %>% setkey(CustomerId)
dtAddr %>% setkey(CustomerId)
dtProd[dtAddr]
# left outer join - swap dt1 with dt2
dtAddr[dtProd, on = "CustomerId"]
data.table의 Merging은 기본모듈인 merge() 과 유사하지만,
dataframe에 기반한 basic merge()는 data.frame내에 변수를 pk로 사용하여 merge하는데 비해
data.table의 Merging은 common key variable 을 pk로 사용하여 merge한다는 점이 다르다.
Example Data
dt1 <- data.table( A=letters[rep(1:3,2)], X=1:6, key="A") dt2 <- data.table( A=letters[rep(2:4,2)], Y=6:1, key="A") dt1 <- data.table( A=letters[rep(1:3,2)], X=1:6) setKey(dt1, "A") dt2 <- data.table( A=letters[rep(2:4,2)], Y=6:1) setKey(dt2, "A")

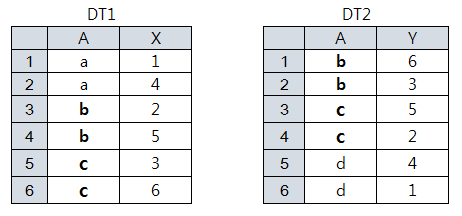
dt1 A X dt2 A Y 1: a 1 1: b 6 2: a 4 2: b 3 3: b 2 3: c 5 4: b 5 4: c 2 5: c 3 5: d 4 6: c 6 6: d 1
all set* functions change their input by reference.
Fast add, remove and update subsets of columns, by reference. :=
Key확인
key(dt1) key(dt2)
[1] "A"
tables() # KEY column reports the key'd columns
NAME NROW NCOL MB COLS KEY [1,] dt1 6 2 1 A,X A [2,] dt2 6 2 1 A,Y A Total: 2MB
update using Two tables
Inner Join
merge(dt1, dt2, by="A") merge(dt1, dt2) # setKey된 A로 merge

A X Y 1: b 2 6 2: b 2 3 3: b 5 6 4: b 5 3 5: c 3 5 6: c 3 2 7: c 6 5 8: c 6 2
Outer join: merge(x = df1, y = df2, by = "CustomerId", all = TRUE)
Left outer: merge(x = df1, y = df2, by = "CustomerId", all.x = TRUE)
Right outer: merge(x = df1, y = df2, by = "CustomerId", all.y = TRUE)
Cross join: merge(x = df1, y = df2, by = NULL)
Subscript Method
A left outer join with df1 on the left using a subscript method would be:
df1[,"State"]<-df2[df1[ ,"Product"], "State"]The other combination of outer joins can be created by mungling the left outer join subscript example. (yeah, I know that's the equivalent of saying "I'll leave it as an exercise for the reader...")
Left (OUTER) JOIN
merge(dt1, dt2, by="A", all.x = T)

A X Y 1: a 1 NA 2: a 4 NA 3: b 2 6 4: b 2 3 5: b 5 6 6: b 5 3 7: c 3 5 8: c 3 2 9: c 6 5 10: c 6 2
Right (OUTER) JOIN
merge(dt1, dt2, by="A", all.y = T) dtResult <- dt1[dt2, on="A"] # [.data.table dtResult <- dt1[dt2, , on="A"] dtResult <- dt1[dt2,.(X,Y), on="A"] dtResult <- dt1[dt2, .(X, i.Y), on="A"] # i.Y = dt2.Y

A X Y 1: b 2 6 2: b 2 3 3: b 5 6 4: b 5 3 5: c 3 5 6: c 3 2 7: c 6 5 8: c 6 2 9: d NA 4 10: d NA 1
Full (OUTER) JOIN
merge(dt1, dt2, all=T)

A X Y 1: a 1 NA 2: a 4 NA 3: b 2 6 4: b 2 3 5: b 5 6 6: b 5 3 7: c 3 5 8: c 3 2 9: c 6 5 10: c 6 2 11: d NA 4 12: d NA 1
예제 Data : 야구 데이터 (from Lahman)
특정 column만 뽑아내기는 data.frame과 data.table이 조금 다르다.
library(Lahman)
data("Pitching")
# Extract specific column
setDF(Pitching)
P0 <- Pitching[ , c('playerID', 'yearID', 'teamID', 'W', 'L', 'G', 'ERA')]
setDT(Pitching)
P1 <- Pitching[ , .(playerID, yearID, teamID, W, L, G, ERA)]
예제 Data : data.table
library(data.table)
set.seed(666)
DT <- data.table( A=rep(c("a","b","c"),each=2), B=c(1:3), C=sample(6), D=sample(6))
setkey(DT, A, B)
setkey는 data.table의 Key로써 A, B를 지정한다. 은 id 변수로 그룹화의 기준으로 생각하면 된다.
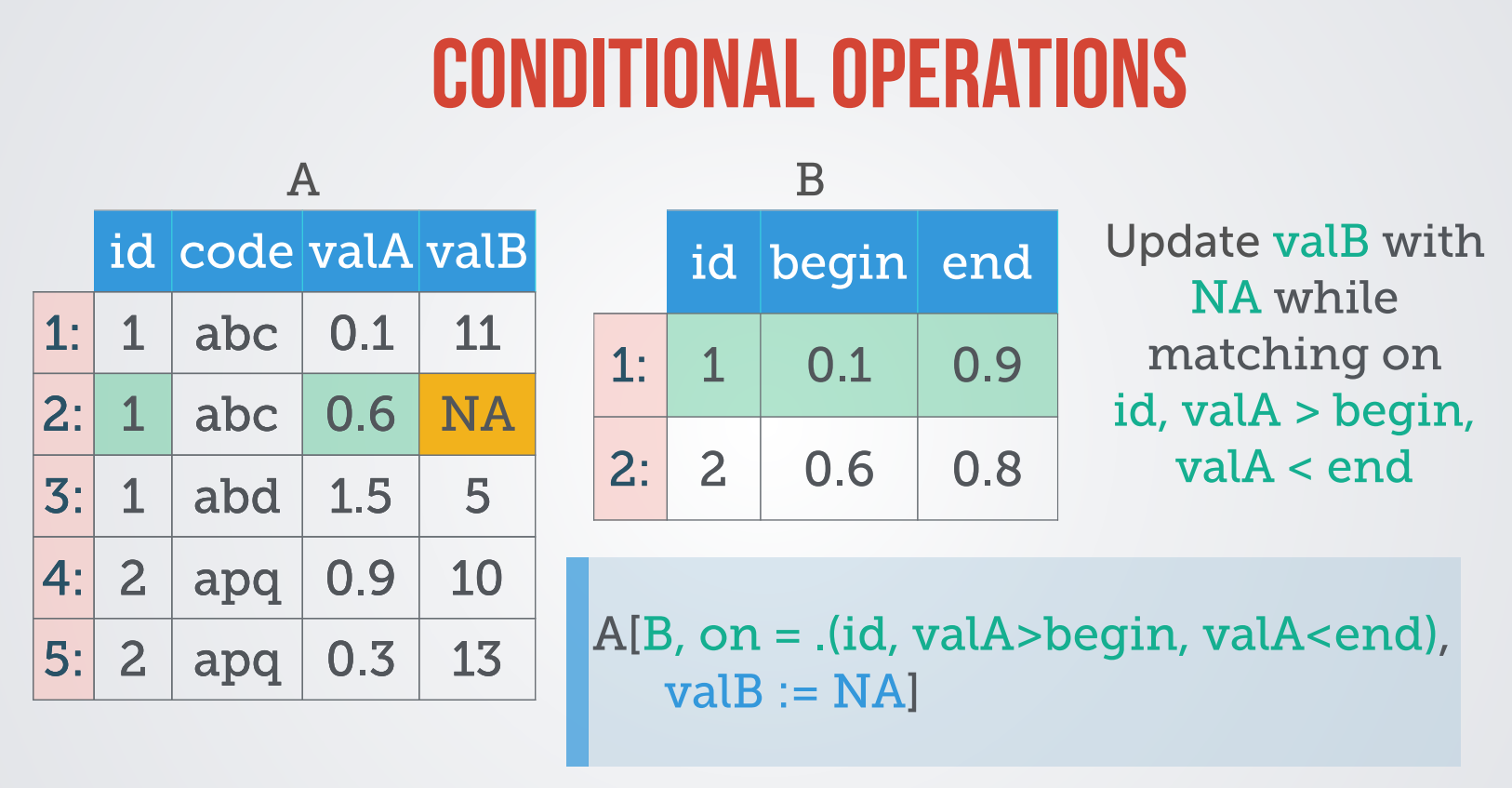
Roll
Join 시 Match되지 않는 데이터 처리
- 기본적으로 NA로 return된다.
- NA로 처리될 row를 지워버릴수 있다.
- Roll을 통해 다른 값을 대체할수 있다.
(LOCF) last observation carried forward => +Inf (or TRUE) 마지막값으로
(NOCB)next observation carried backward => -Inf
rolls the nearest value instead. => "nearest"
DT[.("b", 1:2), on=c("x", "y")] # no match returns NA
x y v
1: b 1 1
2: b 2 NA
DT[.("b", 1:2), on=.(x, y), nomatch=0] # no match row is not returned
x y v
1: b 1 1
DT[.("b", 1:2), on=c("x", "y"), roll=Inf] # locf, nomatch row gets rolled by previous row
x y v
1: b 1 1
2: b 2 1
DT[.("b", 1:2), on=.(x, y), roll=-Inf] # nocb, nomatch row gets rolled by next row
x y v
1: b 1 1
2: b 2 2
. (dot)
가끔 regression formulas에서 볼수 있었던 (ex. lm(y~., data=DT) ) . 는"all other variables"을 의미하지만,
data.table에서는 "list"를 의미한다. any*는 .( ) 를 활용
DT[ , .(x, y)] DT[ , list(x, y)] # identical
.SD (Subset of Data)
.SD는 하나의 data.table의 columns을 각각 sub data.table로 나누어 self reference할수 있게 해준다.
subsetting
: .SD만 단순히 사용하면, 원래 값과 같다.
DT DT[ , .SD] # identical
A B C D 1: a 1 1 3 2: a 2 6 6 3: b 3 5 4 4: b 1 3 2 5: c 2 4 1 6: c 3 2 5
self reference
: A column와 B column을 paste하는 것을 .SD를 활용하여 같은 결과를 구현할수 있다.
paste(DT$A,DT$B,sep="") DT[ , .SD[ , paste(A,B,sep="")], ] # identical
[1] "a1" "a2" "b3" "b1" "c2" "c3
.SDcols
column subsetting
DT[ , .SD, .SDcols=c("B","C","D")]
DT[ , .SD[ ,c("B","C","D")] ] # identical
B C D 1: 1 2 1 2: 2 5 3 3: 3 1 6 4: 1 4 5 5: 2 6 4 6: 3 3 2
ex> Sum all columns
DT[ , lapply(.SD, sum)] # data.table DT[ , colSums(.SD)] # vector
B C D 12 21 21
lapply 활용
.SD는 lapply에서 Column 기준으로 반복
DT[, lapply(.SD, function(x){is.numeric(x)})]
A B C D 1: FALSE TRUE TRUE TRUE
Sum A columns
DT[ , lapply(.SD, sum), .SDcols=2] DT[ , lapply(.SD, sum), .SDcols="B"]
B 1: 12
Sum all columns EXCEPT A
DT[ , lapply(.SD, sum), .SDcols=!"A"]
B C D 1: 12 21 21
grouping BY A => Sum all columns EXCEPT A
DT[ , lapply(.SD, sum), by=A]
A B C D 1: b 4 10 5 2: a 3 6 9 3: c 5 5 7
Sum all columns EXCEPT A, grouping BY B
DT[ , lapply(.SD, sum), by=B, .SDcols=!"A"]
B C D 1: 1 11 10 2: 2 4 5 3: 3 6 6
:= data의 불필요한 Copy를 방지
data.table's reference semantics vignette
Subsetting의 인덱싱 결과는 또 다른 새로운 data.table이다.(즉 원래 DT에 재할당하지 않는 이상, DT는 변하지 않고 유지된다)
따라서, DT의 size가 big한 경우, := 를 사용하여 컬럼을 참조하여 update함으로써 data의 불필요한 Copy를 방지한다.
DT[ , names(DT):= lapply(.SD, as.factor) ]
row selection
sub-data.table의 row index를 활용하여 dynamically하게 원하는 row만 뽑아올수 있다.
아래는 각 그룹의 B열에서 max값을 갖는 row를 뽑아오는 예제.
DT[, .SD[which.max(B)], by=A]
A B C D 1: a 2 4 5 2: b 3 3 3 3: c 3 5 1
DT[, .(max(B)), by=A]
DT[, lapply(.SD, max), .SDcols=c('B'), by=A]
DT[, lapply(.SD, function(x){B=max(x)}), .SDcols=c('B'), by=A]
A B 1: a 2 2: b 3 3: c 3
Convert Column Type
여러 방법을 통해 out_idx를 정의하고, 해당 컬럼에 함수(as.factor)를 적용하여, column type을 covert한다.
data.table이 각 element가 column인 list로 여겨질수 있다는 것이다.
(out_idx)를 괄호로 감싸는 이유는 컬럼명으로 인식하게 하기 위해.
#1. 컬럼 직접 지정
out_idx <- c('A')
out_idx <- c(1)
#2. type확인하여 컬럼 지정
out_idx <- which(sapply(DT, is.character))
#3. colname의 패턴으로 컬럼 지정
out_idx <- grep('A', names(DT))
out_idx <- grep('A', names(DT), value=T) # value=F는 index값
DT[ , (out_idx):=lapply(.SD, as.factor), .SDcols=out_idx]
Data.Frame을 사용해서도 같은 결과를 얻을수 있다.
setDF(DT) # convert to data.frame for illustration sapply(DT[ ,out_idx], is.character)
SD[.N]
.N 은 그룹내의 전체row갯수 . 즉 그룹별로 nrow(.SD)한것과 같다.
ex)
.SD[.N] 은 각 그룹내의 마지막 row값(관측값, obs)을 나타내고, .SD[1L] 은 각 그룹내에 첫번째 관측값을 나타낸다.
DT %>% nrow # 6 DT[ , .SD[.N], ] DT[ , .SD[6], ] DT[ 6, ] DT[, .SD[.N], by=A] DT[c(2,4,6), ]
Performance check
# 10*6 data.table
DT <- replicate(6, sample(seq(100L),10,T)) %>% as.data.table
DT[ , LETTERS[1:2] := .(sample(100L,10,T), sample(100L,10,T))]
# 10000000 * 6 더 큰 데이터테이블 만들기
kk = seq(100L)
nn = 1e7
DT <- replicate(6, sample(kk,nn,T)) %>% as.data.table
DT[ , LETTERS[1:2]:=.(sample(kk,nn,T), sample(kk,nn,T))]
library(microbenchmark)
microbenchmark(times = 100L,
colsums = colSums(DT[ , !c("A", "B"), with = FALSE]),
colsums2 = DT[ , colSums(.SD), .SDcols=!c("A", "B")],
lapplys = DT[ , lapply(.SD, sum), .SDcols = !c("A", "B")])
Unit: milliseconds
expr min lq mean median uq max neval cld
colsums 215.75608 216.39019 222.84491 216.99909 221.86177 395.47297 100 c
colsums2 164.90262 165.44731 166.86141 165.72122 166.05506 181.50616 100 b
lapplys 36.02945 36.82467 37.44593 37.20621 37.45793 48.53117 100 a